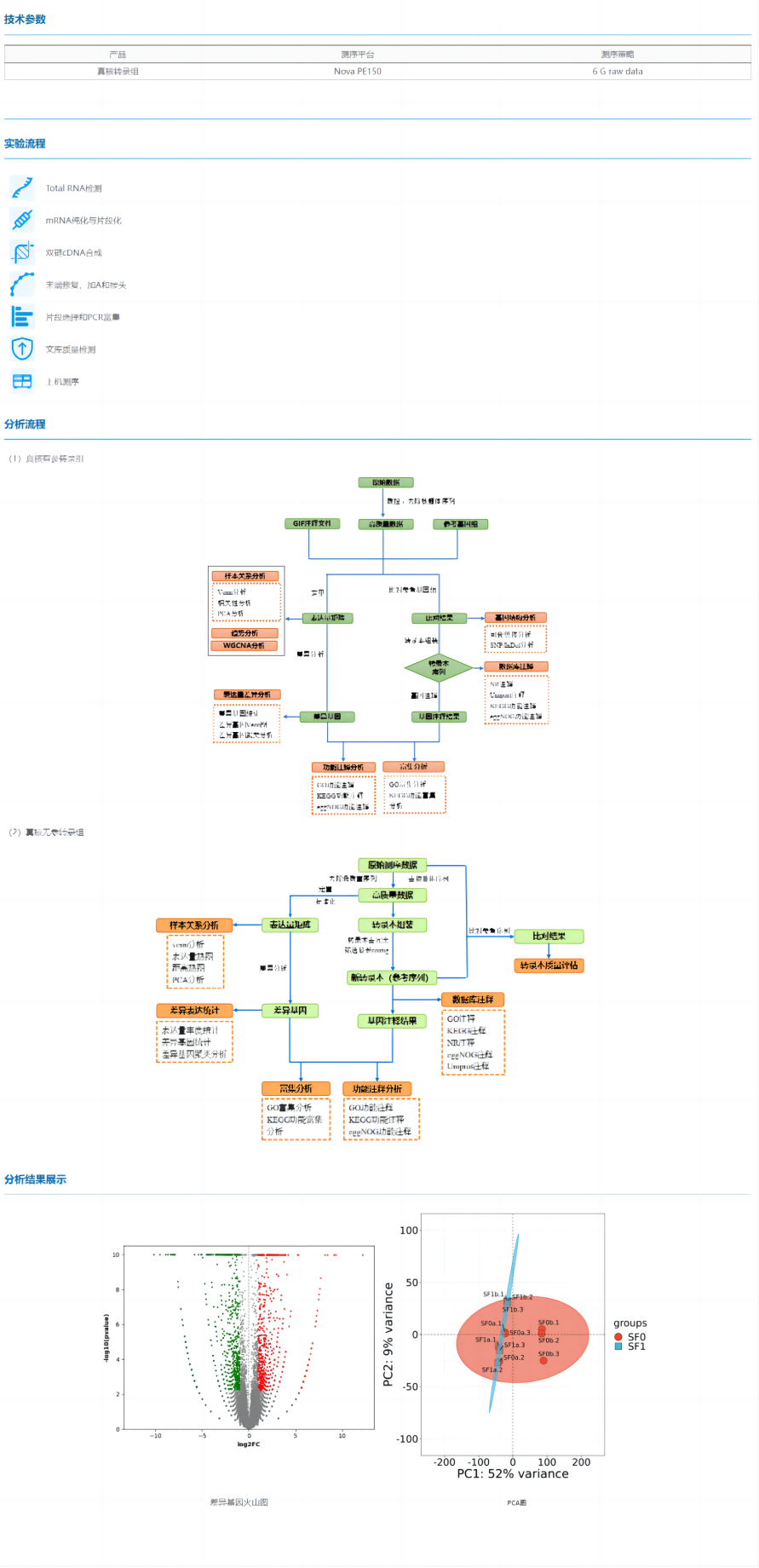
The eukaryotic transcriptome is mainly divided into two categories according to whether there is a reference genome or not, the eukaryotic reference transcriptome and the eukaryotic non-reference transcriptome.
Eukaryotic reference transcriptome: refers to the transcriptome sequencing of eukaryotes or their tissues and cells that have a reference genome. The sequencing results can be compared with the reference genome to verify the sequencing saturation and gene coverage of transcriptome sequencing, and obtain information such as gene structure, gene expression differences, and alternative splicing.
Eukaryotic parameter-free transcriptome: refers to the sequencing of eukaryotic transcriptome in the absence of a reference genome. In eukaryotic non-parametric transcriptome sequencing, firstly, after obtaining the original data, the data needs to be quality controlled and spliced into unigene, and then unigene is used as the reference sequence for subsequent analysis. Differential gene expression analysis and differential gene function enrichment analysis can also be performed on multiple samples to discover functional genes and provide directions for the next step of research.
Features:
l High coverage: comprehensive coding RNA information can be obtained in a single transcriptome sequencing experiment;
l Rich annotation information: through comparison with major databases, relatively complete transcriptome annotation information can be obtained;
l Correlation analysis: Multiple omics data can be combined to carry out cross-omics association analysis.