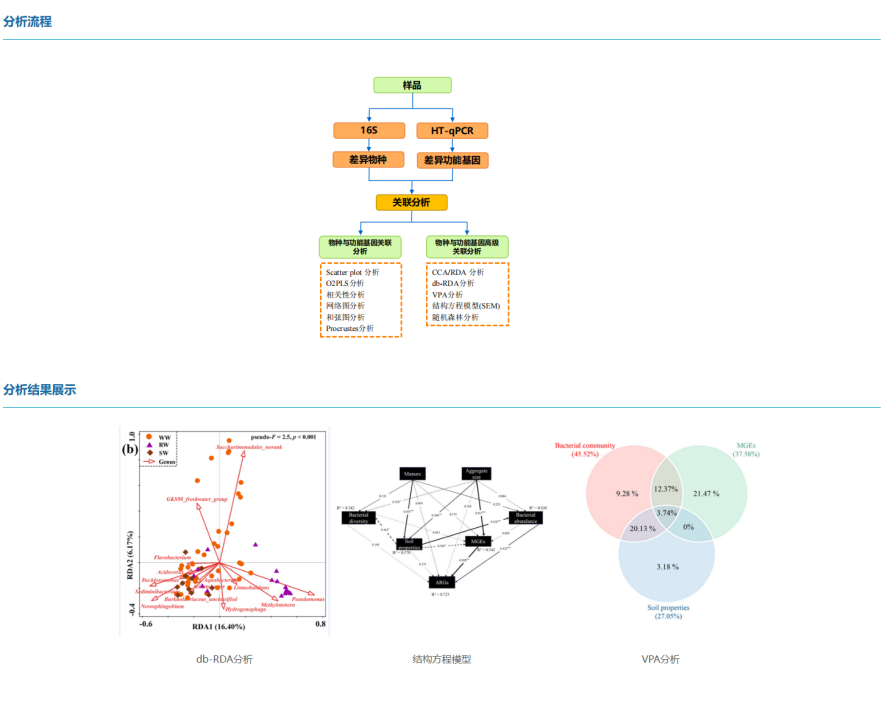
The 16S rDNA gene is a commonly used molecular marker in the phylogenetic classification of prokaryotic microorganisms, and is widely used in the study of microbial ecology. The quantitative method of high-throughput qPCR (HT-qPCR) microarray detection technology has overcome the barriers of sequencing omics methods in terms of co-efficacy and precise quantification, and has been widely used to describe and interpret the functional structure of microorganisms in various ecosystems.
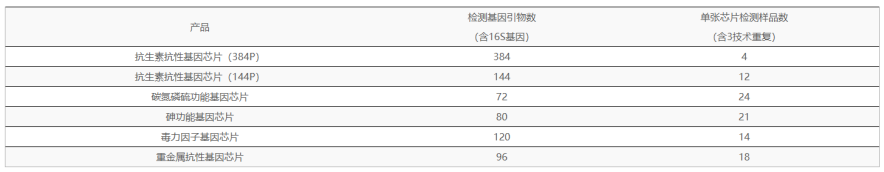
Based on the omics data of 16S and high-throughput qPCR chips, the microbial communities and functional genes with differences between groups were screened out from the level of 16S and functional genes, and then the correlation between the two abundances was jointly analyzed to reveal the correlation between specific microflora and specific functional genes from multiple perspectives, which is of great significance for the study of community structure and function.
Joint analytics benefits
l Mutual verification through omics analysis at the level of microflora and functional genes;
l At the same time, biological problems can be explored from the aspects of "cause" and "effect";
l Explain the association mechanism between microbiota and functional genes, and more comprehensively analyze the species and functional regulation mechanisms in biological processes.